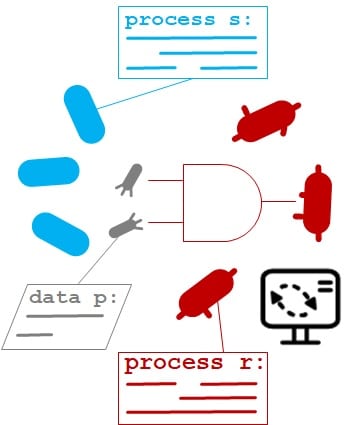
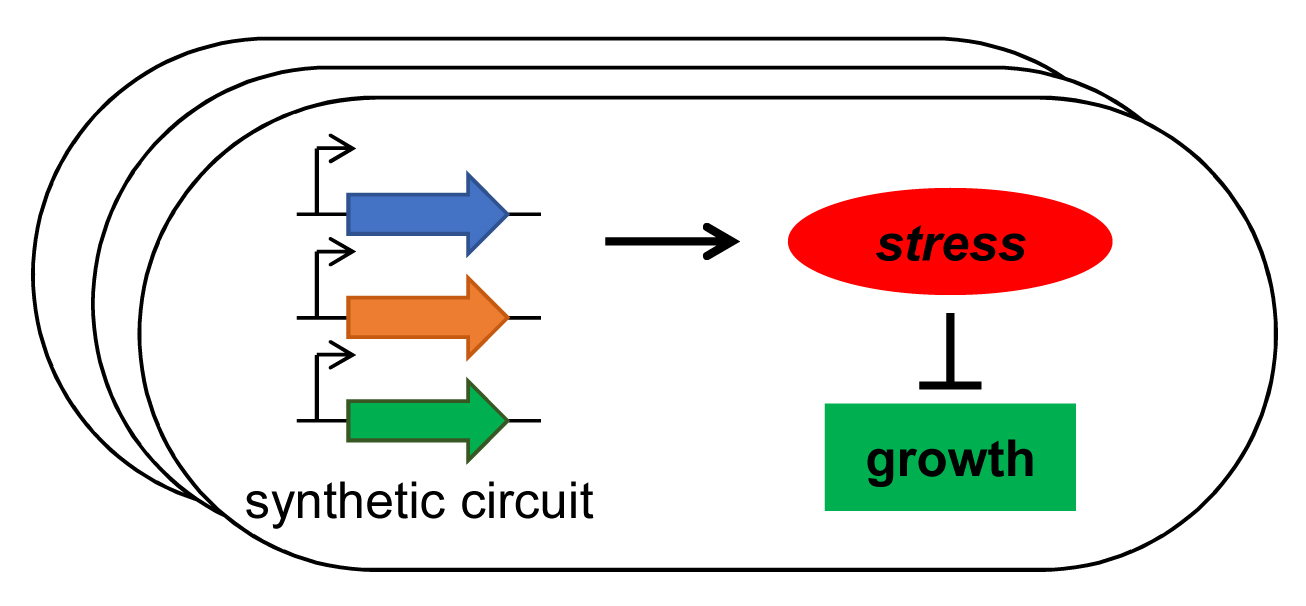
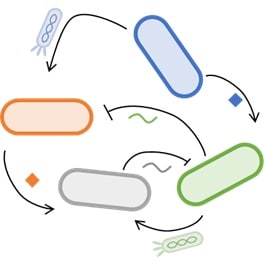
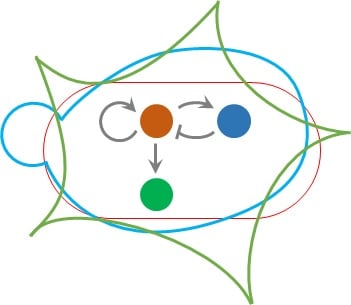
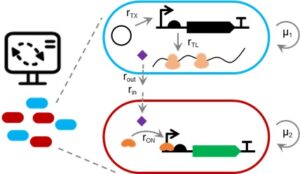
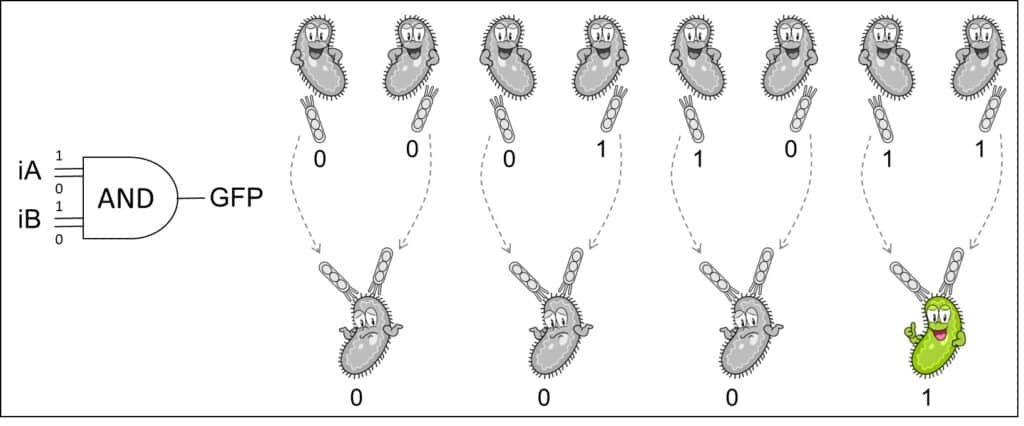
- A. Pujar, A. Pathania, C. Hopper, A. Pandi, C. R. Calderón, M. Függer#, T. Nowak#, M. Kushwaha# (2025). Phage-mediated intercellular CRISPRi for biocomputation in bacterial consortia. Nucleic Acids Research, 53(3), 1–17. DOI: 10.1093/nar/gkae1256
- M.S. Azad*, A.C. Batista*, J.L. Faulon, C.L. Beisel, J. Bonnet, and M. Kushwaha#. (2022). Cell-Free Protein Synthesis from Exonuclease-Deficient Cellular Extracts Utilizing Linear DNA Templates. Journal of Visualized Experiments, 186, 1–13. DOI: 10.3791/64236
- A.C. Batista*, A. Levrier*, P. Soudier*, P. Voyvodic, T. Achmedov, T. Reif-Trauttmansdorff, A. DeVisch, M.C. Gonsaud, J.L. Faulon, C.L. Beisel, J. Bonnet#, and M. Kushwaha#. (2022). Differentially Optimized Cell-Free Buffer Enables Robust Expression from Unprotected Linear DNA in Exonuclease-Deficient Extracts. ACS Synthetic Biology, 11(2), 732–746. DOI: 10.1021/acssynbio.1c00448
- D.-J. Cho, M. Függer, C. Hopper, M. Kushwaha, T. Nowak#, and Q. Soubeyran. (2021). Distributed computation with continual population growth. Distributed Computing, 35(6), 547–569. DOI: 10.1007/s00446-021-00404-8
- A. Pandi*, M. Koch*, P.L. Voyvodic, P. Soudier, J. Bonnet, M. Kushwaha#, and J-L. Faulon#. (2019). Metabolic perceptrons for neural computing in biological systems. Nature Communications, 10(1), 3880. DOI: 10.1038/s41467-019-11889-0
All the team publications are available in the CellComp-MICALIS HAL collection.